 Yes, everybody knows that: we all live in a magnetic field - the Earth. Its field strength is a mere 30-60 micro Tesla (depending on where you live). However, a large metal tube such as a submarine deep in the ocean can cause magnetic disturbances and is therefore detectable. Rumor has it that a certain U-Boat captain founded an NMR company, most likely inspired by his wartime experience evading MADs (magnetic anomaly detectors). A submarine can become electrically charged owing to the difference in the temperature of the upper and lower hull. This is enough to create a difference in salinity of the enclosing sea water, hence a dynamic electric field around "Das Boot". A stealth submarine will have to address this issue obviously. In NMR, shielding high-field magnets is a challenge even now. Incidentally, Mr Tom Clancy has much more on anti-submarine warfare in his classic "The Hunt for Red October" (which most people on Earth have already read):
Yes, everybody knows that: we all live in a magnetic field - the Earth. Its field strength is a mere 30-60 micro Tesla (depending on where you live). However, a large metal tube such as a submarine deep in the ocean can cause magnetic disturbances and is therefore detectable. Rumor has it that a certain U-Boat captain founded an NMR company, most likely inspired by his wartime experience evading MADs (magnetic anomaly detectors). A submarine can become electrically charged owing to the difference in the temperature of the upper and lower hull. This is enough to create a difference in salinity of the enclosing sea water, hence a dynamic electric field around "Das Boot". A stealth submarine will have to address this issue obviously. In NMR, shielding high-field magnets is a challenge even now. Incidentally, Mr Tom Clancy has much more on anti-submarine warfare in his classic "The Hunt for Red October" (which most people on Earth have already read): Back to the issue at hand: At some point in the not so distant future, the full potential of biological NMR will finally be realized. And it will be in the form of 4-dimensional (three spatial and one chemical-shift) NMR spectroscopy. In other words, imaging of, not just the water-protons, but the far more informative metabolites. The limiting factor is, yep, you've guessed it, the magnetic field strength.
Back to the issue at hand: At some point in the not so distant future, the full potential of biological NMR will finally be realized. And it will be in the form of 4-dimensional (three spatial and one chemical-shift) NMR spectroscopy. In other words, imaging of, not just the water-protons, but the far more informative metabolites. The limiting factor is, yep, you've guessed it, the magnetic field strength.Now picture yourself inside a magnet with a set of antennas enclosing your head. And after a certain period of rf irradiation at proton Larmor frequency and FID signal collection and processing, you are ready for the result. And it'll be a spectrum with only a large water peak. That won't be so interesting at all, will it. So, how is an MR image constructed?
Very simple, the FIDs must be encoded in three dimensions. First, along the direction of the north-south direction (the z-axis) of the magnetic field, a field gradient (i.e., the z-gradient) is applied. This gradient causes the Larmor frequency to change linearly, and a slice is selected by choosing the rf frequency corresponding to the Larmor frequency of the center of that slice. For this slice, a preparation gradient, the y-gradient, is then applied for frequency encoding, followed by the x-gradient for phase-encoding. Finally, a 2D Fourier Transform is used to transform the encoded image into the spatial domain. Why the need for a slice? Answer: to make sure enough signals are generated. A single layer of nuclei will have almost negligible signals to work with. [Note: For more information on important aspects of MRI, we recommend this site: http://mritechnicianschools.net].
Present-day clinical MRI is mostly imaging of water-protons. For the ultimate 4-dimensional NMR spectroscopy, an example is shown below:
Again, using an old standby: a rabbit lens (10-mm diameter and 5-mm thickness) incubated for 28 hours in high glucose for maximal sorbitol production. 13C1-glucose was used as the substrate. And NMR experiments were performed using a 9.4-Tesla spectrometer.
Part 1. MR microscopy:
Top: Schematic diagrams of the lens. The two primary regions are the cortex and the nucleus. (Left): section through the anterior-posterior poles and (Right): section through the equatorial plane.
Bottom: A water-proton image, through the equatorial plane, acquired with a spin-echo pulse sequence with TE (echo time)/TR (repetition time) = 3.88/1000ms, a 126x128 matrix, and 32 averages. Image resolution was 40 x 40 x 500 micron.

Part 2. Chemical-shift 13C microscopy:
 At the bottom is a 13C spectrum acquired at 100.61MHz without proton decoupling - hence the multiplets of the biochemicals.
At the bottom is a 13C spectrum acquired at 100.61MHz without proton decoupling - hence the multiplets of the biochemicals.And the top shows color-coded images of glucose (red), sorbitol (yellow), and lactate (purple), respectively. The images were acquired with spin-echo of TE/TR = 9/500ms over 8 hours. A narrow bandwidth selective pulse (4KHz) was used with z-gradient = 800Hz/mm (slice thickness = 5mm). All three metabolites resonated within this bandwidth. The in-plane resolution was 600 x 600 micron. From the images, it is clear that there is a glucose concentration gradient with higher glucose in the cortex. And in contrast, sorbitol appears to re-distribute into the nucleus even though it is produced mostly in the cortex.
Obviously the above still cannot be done on humans. However, 7-Tesla clinical imagers are now coming on line. It is really only a matter of time before the manufacturers start seeking FDA approval for these imagers. For now, an alternative is to collect NMR spectra from, e.g., selected areas of the brain. It is possible to acquire somewhat low-resolution 31P, water-suppressed proton, and even 13C spectra at 1.5 Tesla - the most common magnet field strength used in a clinical setting today.
In the above example, Cho (choline), Cr (creatine), and NAA (N-acetyl aspartate) are clearly seen. Each spectrum is from a volume of interest of 100 x 80 x 15 mm with an acquisition time of 60 min. [Note: A little long, but not intolerable from personal experience. Despite the loud knocking noises from turning gradients, some are known to doze off in the magnet.]
So, here is the current situation: MRI using 1.5-Tesla body scanners has matured as a routine clinical procedure. However, to see the metabolites, the only practical approach is to acquire NMR spectra from a volume of interest in the brain (or other body parts). This is in-vivo NMR spectroscopy, also known as MRS (magnetic resonance spectroscopy). Again, with ultra-high-field clinical magnets now rapidly becoming available, we should expect high-resolution MRS and perhaps even chemical-shift imaging soon.
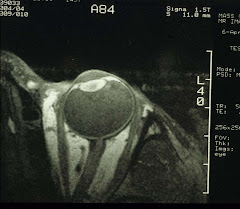
For the eye, we will need to make some adjustment. Instead of the body or the head coil, we use a home-made surface coil that transmits and receives rf. Typically, it is a single-turn 1-inch coil that covers the whole eyeglobe:
 It is then possible to obtain high-resolution MR images of the eye and water-suppressed proton spectra of the vitreous. Again, for the time being.
It is then possible to obtain high-resolution MR images of the eye and water-suppressed proton spectra of the vitreous. Again, for the time being.











1 comment:
Eye doctors routinely use pinhole occluders to determine correctable vision.
The sales of "pinhole glasses", on the other hand, have run into some legal and regulatory problems in the US, chiefly because of unsubstantiated claims such as a "cure" for acquired myopia, etc.
The best way is then for the vendors to sponsor at least a small scale clinical study so that the claims, if any, can be verified.
Examples abound: the testing of progressive bifocals and RGP contacts in slowing the progression of school myopia.
This is the approach that I would advocate, not personal testimonies, however unbiased.
Post a Comment